Cycle fluxes in receptor models#
UNDER CONSTRUCTION#
%%capture
%run receptor_tools.ipynb
As discusssed earlier earlier, the Markov chain tree theorem (also called Hill’s diagrammatic method) provides algebraic expressions for the steady-state probabilities of a receptor model. The function hill_diagramatic_method() below takes a symbolic generator matrix for such a Markov chain as input and, using the Markov chain tree theorem, calculates the steady-state probability of each state.
The receptor models considered here have state-transition diagrams with the topology of a simple symmetric directed graph (connected, no loops). The Markov chain tree theorem always applies to receptor models with these properties. However, there are Markov chains that can not be analyzed in this way (e.g., Markov chains with an infinite number of states).
Receptor model with topology of the House Graph#
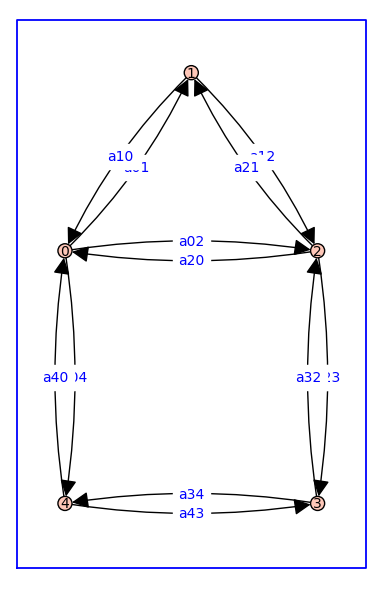
Consider receptor model that has more than one cycle, namely, the house graph.
var('p0 p1 p2 p3 p4 a01 a10 a02 a20 a04 a40 a12 a21 a23 a32 a34 a43')
d = {0: {1:a01, 2:a02, 4:a04}, 1: {0:a10, 2:a12}, 2: {1:a21, 0:a20, 3:a23}, 3: {2:a32, 4:a34}, 4: {3:a43, 0:a40}};
G = DiGraph(d,weighted=True)
vertex_positions = {0: (0, 0), 1: (1, 1.41), 2: (2, 0), 3: (2,-2), 4: (0,-2)}
G.plot(figsize=8,edge_labels=True,pos=vertex_positions,graph_border=True)
Q = generator(G.weighted_adjacency_matrix(sparse=False))
z = hill_diagramatic_method(Q)
for i in range(5):
print('z[%s] ='%i, f'{expand(z[i])}')
z[0] = a10*a20*a32*a40 + a12*a20*a32*a40 + a10*a21*a32*a40 + a10*a20*a34*a40 + a12*a20*a34*a40 + a10*a21*a34*a40 + a10*a23*a34*a40 + a12*a23*a34*a40 + a10*a20*a32*a43 + a12*a20*a32*a43 + a10*a21*a32*a43
z[1] = a01*a20*a32*a40 + a01*a21*a32*a40 + a02*a21*a32*a40 + a01*a20*a34*a40 + a01*a21*a34*a40 + a02*a21*a34*a40 + a01*a23*a34*a40 + a01*a20*a32*a43 + a01*a21*a32*a43 + a02*a21*a32*a43 + a04*a21*a32*a43
z[2] = a02*a10*a32*a40 + a01*a12*a32*a40 + a02*a12*a32*a40 + a02*a10*a34*a40 + a01*a12*a34*a40 + a02*a12*a34*a40 + a02*a10*a32*a43 + a04*a10*a32*a43 + a01*a12*a32*a43 + a02*a12*a32*a43 + a04*a12*a32*a43
z[3] = a02*a10*a23*a40 + a01*a12*a23*a40 + a02*a12*a23*a40 + a04*a10*a20*a43 + a04*a12*a20*a43 + a04*a10*a21*a43 + a02*a10*a23*a43 + a04*a10*a23*a43 + a01*a12*a23*a43 + a02*a12*a23*a43 + a04*a12*a23*a43
z[4] = a04*a10*a20*a32 + a04*a12*a20*a32 + a04*a10*a21*a32 + a04*a10*a20*a34 + a04*a12*a20*a34 + a04*a10*a21*a34 + a02*a10*a23*a34 + a04*a10*a23*a34 + a01*a12*a23*a34 + a02*a12*a23*a34 + a04*a12*a23*a34
In this case, each of the five relative probabilities is a multinomial with 10 terms because there are 10 rooted spanning trees associated to each state (vertex). Each term has 4 factors because any spanning tree of the house graph has 4 edges.
def myHouseGraph():
H = DiGraph()
vertex_vars = ['a0','a1','a2','a3','a4']
edge_vars = ['a','b','c','d','e','f','A','B','C','D','E','F']
V=PolynomialRing(ZZ,names=vertex_vars,order='invlex')
V.inject_variables()
E=PolynomialRing(ZZ,names=edge_vars,order='lex')
E.inject_variables()
H.add_edge(a0,a1,E.gen(edge_vars.index('a')))
H.add_edge(a1,a0,E.gen(edge_vars.index('A')))
H.add_edge(a0,a2,E.gen(edge_vars.index('c')))
H.add_edge(a2,a0,E.gen(edge_vars.index('C')))
H.add_edge(a0,a4,E.gen(edge_vars.index('f')))
H.add_edge(a4,a0,E.gen(edge_vars.index('F')))
H.add_edge(a1,a2,E.gen(edge_vars.index('b')))
H.add_edge(a2,a1,E.gen(edge_vars.index('B')))
H.add_edge(a2,a3,E.gen(edge_vars.index('d')))
H.add_edge(a3,a2,E.gen(edge_vars.index('D')))
H.add_edge(a3,a4,E.gen(edge_vars.index('e')))
H.add_edge(a4,a3,E.gen(edge_vars.index('E')))
return H, E
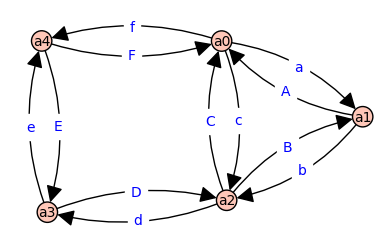
H, P = myHouseGraph()
H.show(edge_labels=True,layout='spring')
Defining a0, a1, a2, a3, a4
Defining a, b, c, d, e, f, A, B, C, D, E, F
def tree_polynomial(G,P=None,root=None):
if P==None:
edge_vars = []
f = 0
#edge_coeff = []
for e in G.edges(sort=True):
f += e[2]
#print(e,e[2].monomials()[0],list(e[2].monomials()[0]))
print(f.variables())
edge_vars = list(f.variables())
# edge_coeff.append(e[2]/e[2].monomials()[0])
P = PolynomialRing(ZZ,names=edge_vars)
Trees = []
if G.is_directed():
if root != None:
T = P(0)
for b in G.in_branchings(source=root,spanning=True):
t = P(1)
for e in b.edges(sort=True):
t *= P(e[2])
T += t
return T
else:
for r in G.vertices(sort=True):
#print('r=',r)
T = P(0)
for b in G.in_branchings(source=r,spanning=True):
#b.show(figsize=10,edge_labels=true)
t = P(1)
for e in b.edges(sort=True):
#print(e)
#t *= P(e[2].monomials()[0])
t *= P(e[2])
T += t
Trees.append(T)
#return Trees
return Trees
else:
T = 0
for b in G.spanning_trees():
t = 1
for e in b.edges(sort=True):
t *= G.edge_label(e[0],e[1])
T += t
return P(T)
TT = tree_polynomial(H,P)
T = tree_polynomial(H,P,a0)
show(T)
for v in H.vertices(sort=True):
T = tree_polynomial(H,P,v)
print(v,':',T)
a0 : b*d*e*F + b*e*C*F + b*C*D*E + b*C*D*F + d*e*A*F + e*A*B*F + e*A*C*F + A*B*D*E + A*B*D*F + A*C*D*E + A*C*D*F
a1 : a*d*e*F + a*e*B*F + a*e*C*F + a*B*D*E + a*B*D*F + a*C*D*E + a*C*D*F + c*e*B*F + c*B*D*E + c*B*D*F + f*B*D*E
a2 : a*b*e*F + a*b*D*E + a*b*D*F + b*c*e*F + b*c*D*E + b*c*D*F + b*f*D*E + c*e*A*F + c*A*D*E + c*A*D*F + f*A*D*E
a3 : a*b*d*E + a*b*d*F + b*c*d*E + b*c*d*F + b*d*f*E + b*f*C*E + c*d*A*E + c*d*A*F + d*f*A*E + f*A*B*E + f*A*C*E
a4 : a*b*d*e + b*c*d*e + b*d*e*f + b*e*f*C + b*f*C*D + c*d*e*A + d*e*f*A + e*f*A*B + e*f*A*C + f*A*B*D + f*A*C*D
def cycle_fluxes(G,P,verbose=True):
Cvert3 = []
Cvert2 = []
for C in G.all_simple_cycles():
if len(C) > 3:
Cvert3.append(C)
else:
Cvert2.append(C)
Cvert2.sort()
Cvert3Sort = Cvert3.copy()
Cvert3Sort.sort()
Cvert3SortP = Cvert3Sort.copy()
for i in range(len(Cvert3Sort)/2):
Cvert3SortP.remove(Cvert3SortP[i][::-1])
Cvert3SortP2 = []
for i in range(3,100): # arbitrary max cycle length
Cvert3SortP2 += [ C for C in Cvert3SortP if len(C)==i ]
Cvert3SortP = Cvert3SortP2.copy()
Cvert3SortM = []
for i in range(len(Cvert3SortP)):
Cvert3SortM.append(Cvert3SortP[i][::-1])
CvertP = Cvert3SortP + Cvert2
CvertM = Cvert3SortM + Cvert2
ncycles = len(CvertP)
CedgeP = []
CedgeM = []
ThetaP = []
ThetaM = []
for i in range(ncycles):
CPe = [ G.edge_label(CvertP[i][j],CvertP[i][j+1]) for j in range(len(CvertP[i])-1)]
CMe = [ G.edge_label(CvertM[i][j],CvertM[i][j+1]) for j in range(len(CvertM[i])-1)]
TPe = prod(CPe)
TMe = prod(CMe)
CedgeP.append(CPe)
ThetaP.append(TPe)
CedgeM.append(CMe)
ThetaM.append(TMe)
Tkappa = []
ThetaPMTkappa = []
for i in range(ncycles):
CP=CvertP[i]
#print('CP=',CP)
H = G.copy()
#show(H,edge_labels=True) ######
H.allow_multiple_edges(True)
#print([None]+CP)
H.merge_vertices([None]+CP) # None becomes vertex 0 (as opposed to a0, a1, ...)
if H.order():
#show(H,edge_labels=True) ######
T = tree_polynomial(H,P,root=0)
else:
T = P(1)
Tkappa.append(T)
ThetaPMTkappa.append((ThetaP[i]-ThetaM[i])*Tkappa[i])
if verbose:
print('\ntotal of',len(Cvert2),'+ 2 *',len(Cvert3)/2,'cycles.\n')
for i in range(ncycles):
H = G.copy()
for e in H.edges(sort=True):
H.set_edge_label(e[0],e[1],0)
for j in range(len(CvertP[i])-1):
H.set_edge_label(CvertP[i][j],CvertP[i][j+1],1)
H.set_edge_label(CvertM[i][j],CvertM[i][j+1],-1)
if ThetaP[i] == ThetaM[i]:
print(i,'=>',CvertP[i],'-',CvertM[i],'=>',CedgeP[i],'-',CedgeM[i],'=>',ThetaP[i]-ThetaM[i],'@ (',Tkappa[i],')')
else:
print(i,'=>',CvertP[i],'-',CvertM[i],'=>',CedgeP[i],'-',CedgeM[i],'=>',(ThetaP[i]-ThetaM[i]).factor(),'@ (',Tkappa[i],')')
H.show(color_by_label=True,edge_labels=True)
return ncycles, CedgeP, CedgeM, ThetaP, ThetaM, Tkappa, ThetaPMTkappa
Trees = tree_polynomial(H)
T = [0]*len(Trees)
for i in range(len(Trees)):
T[i] = Trees[i]
print('T[%s] = ' %(i),T[i])
TT = sum(T)
(a, b, c, d, e, f, A, B, C, D, E, F)
T[0] = A*B*D*E + b*C*D*E + A*C*D*E + b*d*e*F + d*e*A*F + e*A*B*F + b*e*C*F + e*A*C*F + A*B*D*F + b*C*D*F + A*C*D*F
T[1] = a*B*D*E + c*B*D*E + f*B*D*E + a*C*D*E + a*d*e*F + a*e*B*F + c*e*B*F + a*e*C*F + a*B*D*F + c*B*D*F + a*C*D*F
T[2] = a*b*D*E + b*c*D*E + b*f*D*E + c*A*D*E + f*A*D*E + a*b*e*F + b*c*e*F + c*e*A*F + a*b*D*F + b*c*D*F + c*A*D*F
T[3] = a*b*d*E + b*c*d*E + b*d*f*E + c*d*A*E + d*f*A*E + f*A*B*E + b*f*C*E + f*A*C*E + a*b*d*F + b*c*d*F + c*d*A*F
T[4] = a*b*d*e + b*c*d*e + b*d*e*f + c*d*e*A + d*e*f*A + e*f*A*B + b*e*f*C + e*f*A*C + f*A*B*D + b*f*C*D + f*A*C*D
ncycles, CedgeP, CedgeM, ThetaP, ThetaM, Tkappa, ThetaPMTkappa = cycle_fluxes(H,P,verbose=True)
for i in range(ncycles):
print('(',ThetaP[i],'-',ThetaM[i],') @ (',Tkappa[i],')')
total of 6 + 2 * 3 cycles.
0 => [a0, a1, a2, a0] - [a0, a2, a1, a0] => [a, b, C] - [c, B, A] => (-1) * (-a*b*C + c*A*B) @ ( e*F + D*E + D*F )
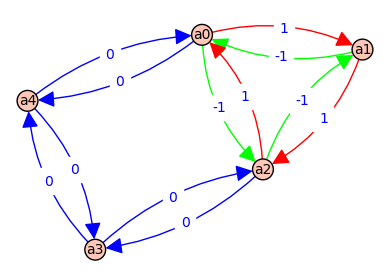
1 => [a0, a2, a3, a4, a0] - [a0, a4, a3, a2, a0] => [c, d, e, F] - [f, E, D, C] => (-1) * (-c*d*e*F + f*C*D*E) @ ( b + A )
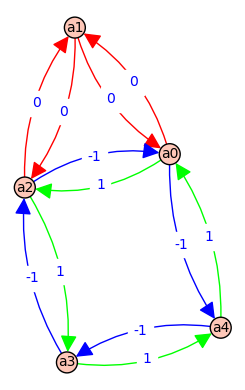
2 => [a0, a1, a2, a3, a4, a0] - [a0, a4, a3, a2, a1, a0] => [a, b, d, e, F] - [f, E, D, B, A] => (-1) * (-a*b*d*e*F + f*A*B*D*E) @ ( 1 )
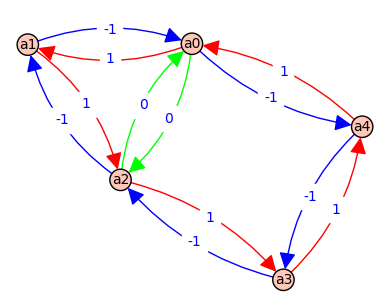
3 => [a0, a1, a0] - [a0, a1, a0] => [a, A] - [a, A] => 0 @ ( d*e*F + e*B*F + e*C*F + B*D*E + B*D*F + C*D*E + C*D*F )
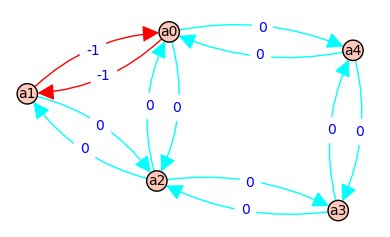
4 => [a0, a2, a0] - [a0, a2, a0] => [c, C] - [c, C] => 0 @ ( b*e*F + b*D*E + b*D*F + e*A*F + A*D*E + A*D*F )
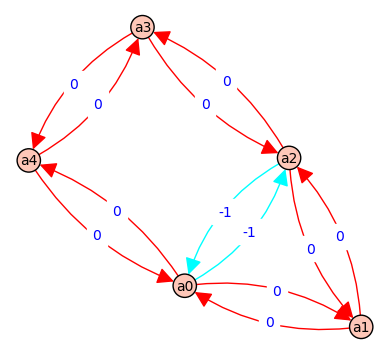
5 => [a0, a4, a0] - [a0, a4, a0] => [f, F] - [f, F] => 0 @ ( b*d*e + b*e*C + b*C*D + d*e*A + e*A*B + e*A*C + A*B*D + A*C*D )
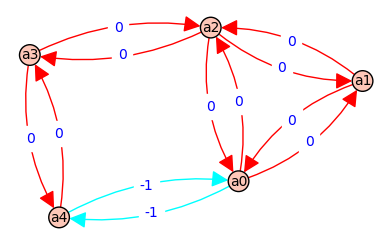
6 => [a1, a2, a1] - [a1, a2, a1] => [b, B] - [b, B] => 0 @ ( a*e*F + a*D*E + a*D*F + c*e*F + c*D*E + c*D*F + f*D*E )
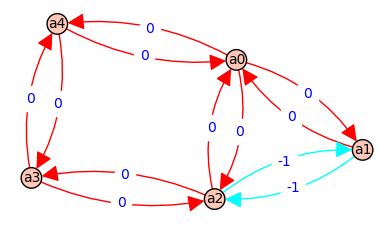
7 => [a2, a3, a2] - [a2, a3, a2] => [d, D] - [d, D] => 0 @ ( a*b*E + a*b*F + b*c*E + b*c*F + b*f*E + c*A*E + c*A*F + f*A*E )
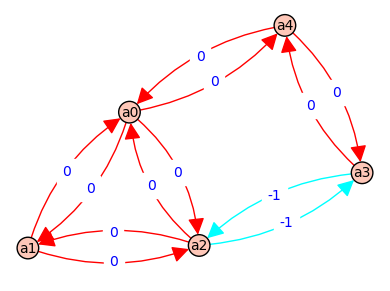
8 => [a4, a3, a4] - [a4, a3, a4] => [E, e] - [E, e] => 0 @ ( a*b*d + b*c*d + b*d*f + b*f*C + c*d*A + d*f*A + f*A*B + f*A*C )
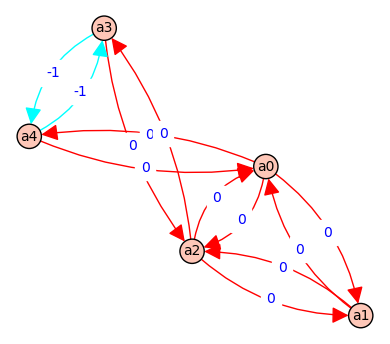
( a*b*C - c*A*B ) @ ( e*F + D*E + D*F )
( c*d*e*F - f*C*D*E ) @ ( b + A )
( a*b*d*e*F - f*A*B*D*E ) @ ( 1 )
( a*A - a*A ) @ ( d*e*F + e*B*F + e*C*F + B*D*E + B*D*F + C*D*E + C*D*F )
( c*C - c*C ) @ ( b*e*F + b*D*E + b*D*F + e*A*F + A*D*E + A*D*F )
( f*F - f*F ) @ ( b*d*e + b*e*C + b*C*D + d*e*A + e*A*B + e*A*C + A*B*D + A*C*D )
( b*B - b*B ) @ ( a*e*F + a*D*E + a*D*F + c*e*F + c*D*E + c*D*F + f*D*E )
( d*D - d*D ) @ ( a*b*E + a*b*F + b*c*E + b*c*F + b*f*E + c*A*E + c*A*F + f*A*E )
( e*E - e*E ) @ ( a*b*d + b*c*d + b*d*f + b*f*C + c*d*A + d*f*A + f*A*B + f*A*C )

