Markov chain receptor models#
Symbolic vertex and edge labels#
Recall the three state model discussed above (Ligands). Using generic notation for the path graph on three vertices,
var('p0 p1 p2 a01 a10 a12 a21')
G=graphs.PathGraph(3).to_directed()
G.relabel({0:p0,1:p1,2:p2})
G.set_edge_label(p0,p1,a01)
G.set_edge_label(p1,p0,a10)
G.set_edge_label(p1,p2,a12)
G.set_edge_label(p2,p1,a21)
G.show(figsize=4,edge_labels=True)
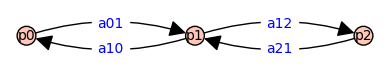
We assign symbolic variables to the vertices and edges of \(G\) because this allows us to produce symbolic expressions important quantities using methods available in Sagemath’s module for graphs and digraphs. For example, the weighted adjacency matrix associated with graph $G$ above is
A = G.weighted_adjacency_matrix()
The combinatorial Laplacian matrix is \(L=D-A\) where \(D\) is a diagonal matrix given by the column sum of \(A\).
L = diagonal_matrix(sum(A.T))-A
show(L)
Note
In the code block above, I prefer to write sum(A.T) rather than sum(A.columns()), but these are equal.
sum(A.T) == sum(A.columns())
True
Generator matrix of Markov chain#
The Markov chain with states and transistions of \(G\) has a generator matrix \(Q\) that is the opposite (additive inverse) of the Laplacian matrix (\(L\)). That is, \(Q=-L=A-D\). We define a function that calculates the symbolic generator matrix $Q$ from the weighted adjacency matrix \(A\).
def generator(A):
return A-diagonal_matrix(sum(A.T))
Calling the function generator() gives the result we expect:
Q = generator(A)
show(Q)
Note that the sums of the columns are zero, reflecting the conservation of probability.
sum(Q.columns()) == sum(Q.T)
True
Markov chain tree theorem#
Using the Markov chain tree theorem, it is straightforward to find symbolic expressions for the steady-state probabilities of each receptor state (p0, p1, p2). The relative probabilities are given by the sum of the weights of all the spanning trees rooted in vertex i. Because this three-state receptor model has no cycles, there is only one directed spanning tree rooted in each vertex (shown below).
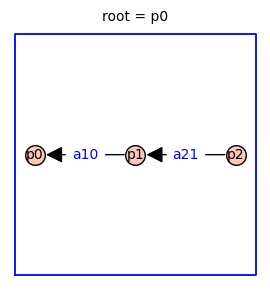
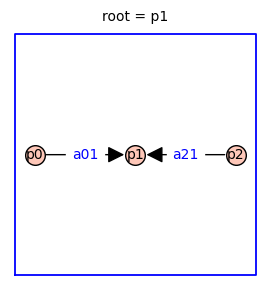
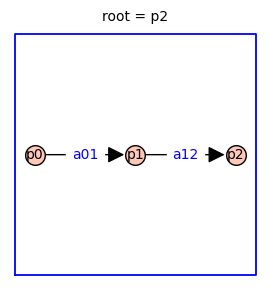
The relative probability of state i is given by the determinant of the Laplacian matrix with row i and column i removed.
In this case, each relative probability is a monomial with two factors.
z0 = Q[[1,2],[1,2]].determinant().simplify_full()
z1 = Q[[0,2],[0,2]].determinant().simplify_full()
z2 = Q[[0,1],[0,1]].determinant().simplify_full()
print(f'[ z0 : z1 : z2 ] = [ {z0} : {z1} : {z2} ]')
[ z0 : z1 : z2 ] = [ a10*a21 : a01*a21 : a01*a12 ]
The normalized probabilities are:
zT = z0+z1+z2
p0 = z0/zT
p1 = z1/zT
p2 = z2/zT
show(table([[f'{p0=}'],[f'{p1=}'],[f'{p2=}']]))
| p0=a10*a21/(a01*a12 + a01*a21 + a10*a21) |
| p1=a01*a21/(a01*a12 + a01*a21 + a10*a21) |
| p2=a01*a12/(a01*a12 + a01*a21 + a10*a21) |
This three-state receptor model has no cycles. For this reason the steady-state probabilities will satisfy detailed balance. This implies that the steady-state probability distribution can be written in terms of equilibrium constants (as opposed to rate constants). Define equilibrium constants as \(\kappa_{j}=a_{ij}/a_{ji}\) for \(i < j\) whenever vertex \(i\) and \(j\) are adjacent. Dividing the numerator and denominator of each \(p_i\) by \(a_{10}a_{21}\) yields
This confirms that the steady-state probabilities of this three-state receptor model can be written in terms of equilibrium constants.

