Enumerating allosteric parameters#
The notebook titled receptor_tools.ipynb includes the function enumerate_allosteric_parameters(G) which takes a graph G and returns the \(\kappa\) and \(\eta\) values for a dimer model with the topology of the reduced graph power \(G^{(2)}\). The default method uses \(e_1, e_2, \ldots , e_v\) while method='alpha' gives single digit edge labels (\(b,c,\ldots\)).
%%capture
%run receptor_tools.ipynb
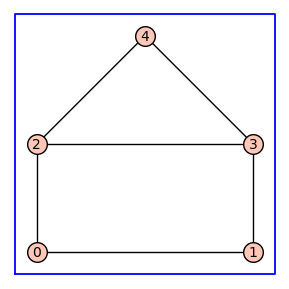
Below we will consider a receptor model with the topology of the house graph. To begin, we construct the graph G with integer vertices. This graph is the input argument for enumerate_allosteric_parameters(G), which returns the 4-tuple (H, T, KappaEta, A). The following code blocks describe each element of this tuple of outputs.
G=graphs.HouseGraph()
G.show(figsize=4,graph_border=True)
(H, T, KappaEta, A) = enumerate_allosteric_parameters(G,method='alpha')
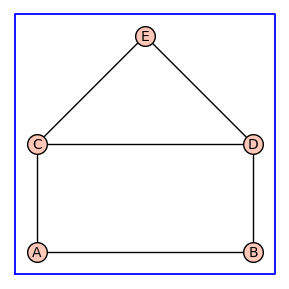
The method = alpha in enumerate_allosteric_parameters(G) evokes the construction of the corresponding graph with monomial vertices. The resulting graph is returned as first element of a 4-tuple output, which in this case is H.
H.show(figsize=4,graph_border=True)
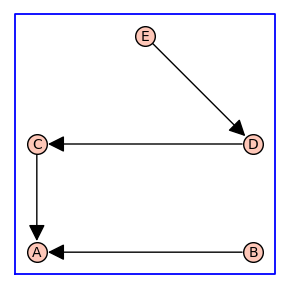
The 2nd element of the 4-tuple output is a spanning tree T of G constructed by enumerate_allosteric_parameters() so that the vertex labels are consistent with a breadth first traversal.
T.show(figsize=4,graph_border=True)
The 3rd element of the output, KappaEta, is a list of lists of the relative probability of each state.
KappaEta
[[1, 2*kappa_b, 2*kappa_c, 2*kappa_c*kappa_d, 2*kappa_c*kappa_d*kappa_e],
[0,
kappa_b^2*eta_bb,
2*kappa_b*kappa_c*eta_bc,
2*kappa_b*kappa_c*kappa_d*eta_bc*eta_bd,
2*kappa_b*kappa_c*kappa_d*kappa_e*eta_bc*eta_bd*eta_be],
[0,
0,
kappa_c^2*eta_cc,
2*kappa_c^2*kappa_d*eta_cc*eta_cd,
2*kappa_c^2*kappa_d*kappa_e*eta_cc*eta_cd*eta_ce],
[0,
0,
0,
kappa_c^2*kappa_d^2*eta_cc*eta_cd^2*eta_dd,
2*kappa_c^2*kappa_d^2*kappa_e*eta_cc*eta_cd^2*eta_dd*eta_ce*eta_de],
[0,
0,
0,
0,
kappa_c^2*kappa_d^2*kappa_e^2*eta_cc*eta_cd^2*eta_dd*eta_ce^2*eta_de^2*eta_ee]]
The command show(table(KappaEta)) displays these symbolic expressions as a table with LaTeX formated entries.
show(table(KappaEta))
| \(1\) | \(2 \kappa_{\mathit{b}}\) | \(2 \kappa_{\mathit{c}}\) | \(2 \kappa_{\mathit{c}} \kappa_{\mathit{d}}\) | \(2 \kappa_{\mathit{c}} \kappa_{\mathit{d}} \kappa_{\mathit{e}}\) |
| \(0\) | \(\kappa_{\mathit{b}}^{2} \eta_{\mathit{bb}}\) | \(2 \kappa_{\mathit{b}} \kappa_{\mathit{c}} \eta_{\mathit{bc}}\) | \(2 \kappa_{\mathit{b}} \kappa_{\mathit{c}} \kappa_{\mathit{d}} \eta_{\mathit{bc}} \eta_{\mathit{bd}}\) | \(2 \kappa_{\mathit{b}} \kappa_{\mathit{c}} \kappa_{\mathit{d}} \kappa_{\mathit{e}} \eta_{\mathit{bc}} \eta_{\mathit{bd}} \eta_{\mathit{be}}\) |
| \(0\) | \(0\) | \(\kappa_{\mathit{c}}^{2} \eta_{\mathit{cc}}\) | \(2 \kappa_{\mathit{c}}^{2} \kappa_{\mathit{d}} \eta_{\mathit{cc}} \eta_{\mathit{cd}}\) | \(2 \kappa_{\mathit{c}}^{2} \kappa_{\mathit{d}} \kappa_{\mathit{e}} \eta_{\mathit{cc}} \eta_{\mathit{cd}} \eta_{\mathit{ce}}\) |
| \(0\) | \(0\) | \(0\) | \(\kappa_{\mathit{c}}^{2} \kappa_{\mathit{d}}^{2} \eta_{\mathit{cc}} \eta_{\mathit{cd}}^{2} \eta_{\mathit{dd}}\) | \(2 \kappa_{\mathit{c}}^{2} \kappa_{\mathit{d}}^{2} \kappa_{\mathit{e}} \eta_{\mathit{cc}} \eta_{\mathit{cd}}^{2} \eta_{\mathit{dd}} \eta_{\mathit{ce}} \eta_{\mathit{de}}\) |
| \(0\) | \(0\) | \(0\) | \(0\) | \(\kappa_{\mathit{c}}^{2} \kappa_{\mathit{d}}^{2} \kappa_{\mathit{e}}^{2} \eta_{\mathit{cc}} \eta_{\mathit{cd}}^{2} \eta_{\mathit{dd}} \eta_{\mathit{ce}}^{2} \eta_{\mathit{de}}^{2} \eta_{\mathit{ee}}\) |
The 4th element of the output is the relevant polynomial ring where the indeterminants are the \(\kappa_i\) and \(\eta_{ij}\) as symbolic variables for \(0 \leq i,j \leq v-1\).
show(A)
The cubic ternary complex model#
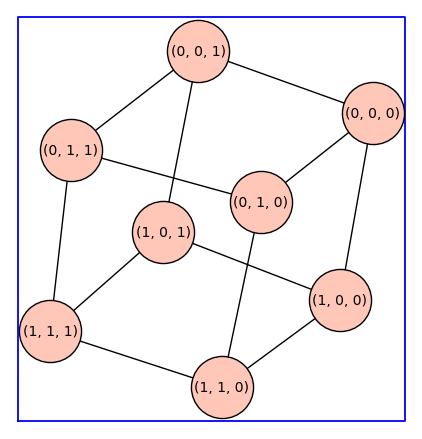
The next example is reminicent of the cubic ternary complex model. Note that graphs.GridGraph([2,2,2]) has tuples as vertices. We use cannonical_label() to make the vertices integers. This is necessary because enumerate_allosteric_parameters() requires a graph G with integer vertices.
G=graphs.GridGraph([2,2,2])
G.show(figsize=6,graph_border=True,vertex_size=2000)
G=G.canonical_label()
G.show(figsize=4,graph_border=True)
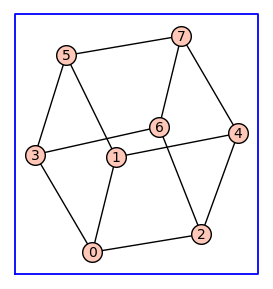
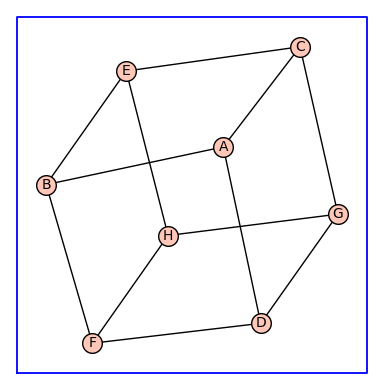
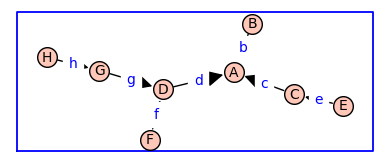
Using the cube \(Q_3\) as the topology of a receptor monomer, enumerate_allosteric_parameters() enumerates the allosteric parameters for the receptor dimer with topology \(Q_3^{(2)}\). The argument show=True in enumerate_allosteric_parameters() causes the display of H, Tand KappaEta.
(H, T, KappaEta, A) = enumerate_allosteric_parameters(G,method='alpha',show=True)
| \(1\) | \(2 \kappa_{\mathit{b}}\) | \(2 \kappa_{\mathit{c}}\) | \(2 \kappa_{\mathit{d}}\) | \(2 \kappa_{\mathit{c}} \kappa_{\mathit{e}}\) | \(2 \kappa_{\mathit{d}} \kappa_{\mathit{f}}\) | \(2 \kappa_{\mathit{d}} \kappa_{\mathit{g}}\) | \(2 \kappa_{\mathit{d}} \kappa_{\mathit{g}} \kappa_{\mathit{h}}\) |
| \(0\) | \(\kappa_{\mathit{b}}^{2} \eta_{\mathit{bb}}\) | \(2 \kappa_{\mathit{b}} \kappa_{\mathit{c}} \eta_{\mathit{bc}}\) | \(2 \kappa_{\mathit{b}} \kappa_{\mathit{d}} \eta_{\mathit{bd}}\) | \(2 \kappa_{\mathit{b}} \kappa_{\mathit{c}} \kappa_{\mathit{e}} \eta_{\mathit{bc}} \eta_{\mathit{be}}\) | \(2 \kappa_{\mathit{b}} \kappa_{\mathit{d}} \kappa_{\mathit{f}} \eta_{\mathit{bd}} \eta_{\mathit{bf}}\) | \(2 \kappa_{\mathit{b}} \kappa_{\mathit{d}} \kappa_{\mathit{g}} \eta_{\mathit{bd}} \eta_{\mathit{bg}}\) | \(2 \kappa_{\mathit{b}} \kappa_{\mathit{d}} \kappa_{\mathit{g}} \kappa_{\mathit{h}} \eta_{\mathit{bd}} \eta_{\mathit{bg}} \eta_{\mathit{bh}}\) |
| \(0\) | \(0\) | \(\kappa_{\mathit{c}}^{2} \eta_{\mathit{cc}}\) | \(2 \kappa_{\mathit{c}} \kappa_{\mathit{d}} \eta_{\mathit{cd}}\) | \(2 \kappa_{\mathit{c}}^{2} \kappa_{\mathit{e}} \eta_{\mathit{cc}} \eta_{\mathit{ce}}\) | \(2 \kappa_{\mathit{c}} \kappa_{\mathit{d}} \kappa_{\mathit{f}} \eta_{\mathit{cd}} \eta_{\mathit{cf}}\) | \(2 \kappa_{\mathit{c}} \kappa_{\mathit{d}} \kappa_{\mathit{g}} \eta_{\mathit{cd}} \eta_{\mathit{cg}}\) | \(2 \kappa_{\mathit{c}} \kappa_{\mathit{d}} \kappa_{\mathit{g}} \kappa_{\mathit{h}} \eta_{\mathit{cd}} \eta_{\mathit{cg}} \eta_{\mathit{ch}}\) |
| \(0\) | \(0\) | \(0\) | \(\kappa_{\mathit{d}}^{2} \eta_{\mathit{dd}}\) | \(2 \kappa_{\mathit{c}} \kappa_{\mathit{d}} \kappa_{\mathit{e}} \eta_{\mathit{cd}} \eta_{\mathit{de}}\) | \(2 \kappa_{\mathit{d}}^{2} \kappa_{\mathit{f}} \eta_{\mathit{dd}} \eta_{\mathit{df}}\) | \(2 \kappa_{\mathit{d}}^{2} \kappa_{\mathit{g}} \eta_{\mathit{dd}} \eta_{\mathit{dg}}\) | \(2 \kappa_{\mathit{d}}^{2} \kappa_{\mathit{g}} \kappa_{\mathit{h}} \eta_{\mathit{dd}} \eta_{\mathit{dg}} \eta_{\mathit{dh}}\) |
| \(0\) | \(0\) | \(0\) | \(0\) | \(\kappa_{\mathit{c}}^{2} \kappa_{\mathit{e}}^{2} \eta_{\mathit{cc}} \eta_{\mathit{ce}}^{2} \eta_{\mathit{ee}}\) | \(2 \kappa_{\mathit{c}} \kappa_{\mathit{d}} \kappa_{\mathit{e}} \kappa_{\mathit{f}} \eta_{\mathit{cd}} \eta_{\mathit{de}} \eta_{\mathit{cf}} \eta_{\mathit{ef}}\) | \(2 \kappa_{\mathit{c}} \kappa_{\mathit{d}} \kappa_{\mathit{e}} \kappa_{\mathit{g}} \eta_{\mathit{cd}} \eta_{\mathit{de}} \eta_{\mathit{cg}} \eta_{\mathit{eg}}\) | \(2 \kappa_{\mathit{c}} \kappa_{\mathit{d}} \kappa_{\mathit{e}} \kappa_{\mathit{g}} \kappa_{\mathit{h}} \eta_{\mathit{cd}} \eta_{\mathit{de}} \eta_{\mathit{cg}} \eta_{\mathit{eg}} \eta_{\mathit{ch}} \eta_{\mathit{eh}}\) |
| \(0\) | \(0\) | \(0\) | \(0\) | \(0\) | \(\kappa_{\mathit{d}}^{2} \kappa_{\mathit{f}}^{2} \eta_{\mathit{dd}} \eta_{\mathit{df}}^{2} \eta_{\mathit{ff}}\) | \(2 \kappa_{\mathit{d}}^{2} \kappa_{\mathit{f}} \kappa_{\mathit{g}} \eta_{\mathit{dd}} \eta_{\mathit{df}} \eta_{\mathit{dg}} \eta_{\mathit{fg}}\) | \(2 \kappa_{\mathit{d}}^{2} \kappa_{\mathit{f}} \kappa_{\mathit{g}} \kappa_{\mathit{h}} \eta_{\mathit{dd}} \eta_{\mathit{df}} \eta_{\mathit{dg}} \eta_{\mathit{fg}} \eta_{\mathit{dh}} \eta_{\mathit{fh}}\) |
| \(0\) | \(0\) | \(0\) | \(0\) | \(0\) | \(0\) | \(\kappa_{\mathit{d}}^{2} \kappa_{\mathit{g}}^{2} \eta_{\mathit{dd}} \eta_{\mathit{dg}}^{2} \eta_{\mathit{gg}}\) | \(2 \kappa_{\mathit{d}}^{2} \kappa_{\mathit{g}}^{2} \kappa_{\mathit{h}} \eta_{\mathit{dd}} \eta_{\mathit{dg}}^{2} \eta_{\mathit{gg}} \eta_{\mathit{dh}} \eta_{\mathit{gh}}\) |
| \(0\) | \(0\) | \(0\) | \(0\) | \(0\) | \(0\) | \(0\) | \(\kappa_{\mathit{d}}^{2} \kappa_{\mathit{g}}^{2} \kappa_{\mathit{h}}^{2} \eta_{\mathit{dd}} \eta_{\mathit{dg}}^{2} \eta_{\mathit{gg}} \eta_{\mathit{dh}}^{2} \eta_{\mathit{gh}}^{2} \eta_{\mathit{hh}}\) |
Larger example#
We conclude with an example of a receptor model with more states. The topology used for the monomer model is given by the Petersen graph. Here we use the method integer rather than alpha. Using show=False suppresses the output. One can still work with the ring A, show the BFS spanning tree, and list the probability of each state.
(G, T, KappaEta, A) = enumerate_allosteric_parameters(graphs.PetersenGraph(),method='integer',show=False)
A.inject_variables()
Defining kappa_1, kappa_2, kappa_3, kappa_4, kappa_5, kappa_6, kappa_7, kappa_8, kappa_9, eta_11, eta_12, eta_22, eta_13, eta_23, eta_33, eta_14, eta_24, eta_34, eta_44, eta_15, eta_25, eta_35, eta_45, eta_55, eta_16, eta_26, eta_36, eta_46, eta_56, eta_66, eta_17, eta_27, eta_37, eta_47, eta_57, eta_67, eta_77, eta_18, eta_28, eta_38, eta_48, eta_58, eta_68, eta_78, eta_88, eta_19, eta_29, eta_39, eta_49, eta_59, eta_69, eta_79, eta_89, eta_99
T.show(figsize=6,graph_border=True,edge_labels=True)
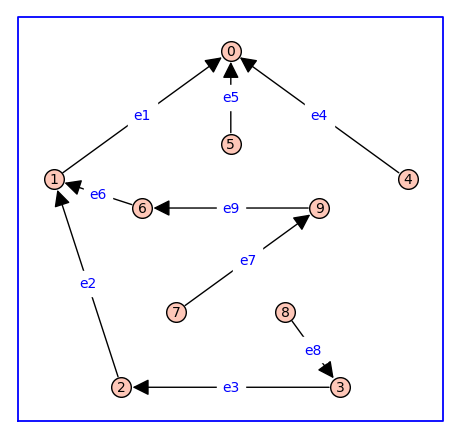
The code block below shows how one can list the symbolic expressions for the relative probabilities of each state in the model receptor dimer.
from IPython.display import display, Math
for i,k in enumerate(flatten(KappaEta)):
if k != 0:
display(Math(latex(i)+':'+latex(k)))

