Tools and Examples - Part 2#
Reduced graph powers#
The notebook titled receptor_tools.ipynb (see Appendix) contains function definitions that will prove useful as we further explore receptor modeling. The command %run receptor_tools.ipynb loads these function definitions, and the remainder of this notebook illustrates how to use some of them. The focus here is on tools that faciliate constructoin of graph powers and reduced graph powers.
%%capture
%run receptor_tools.ipynb
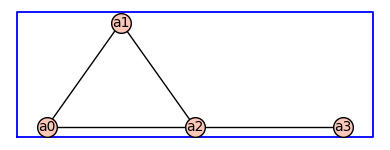
To begin, we specify the states and transitions of a receptor model as an undirected graph. As an example, will use the following four-state model.
pos = {0: (0, 0), 1: (1, 1.41), 2: (2, 0), 3: (4,0)} # vertex positions
G0 = Graph({0: [1, 2], 1: [2], 2: [3]},pos=pos)
G = add_vertex_monomials(G0)
G.show(figsize=4,graph_border=True)
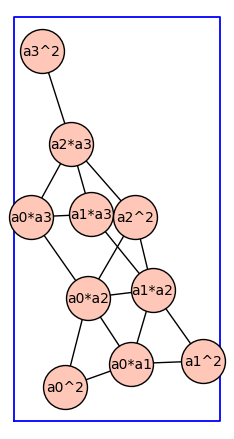
Next we produce the reduced graph power \(G^{(2)}\). Each vertex is a monomial where the indeterminants are given by the vertices of {math}G`.
G2 = reduced_cartesian_power_ver2(G,2)
G2.show(figsize=6,graph_border=True,vertex_size=1000)
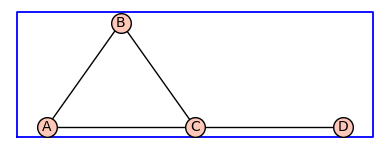
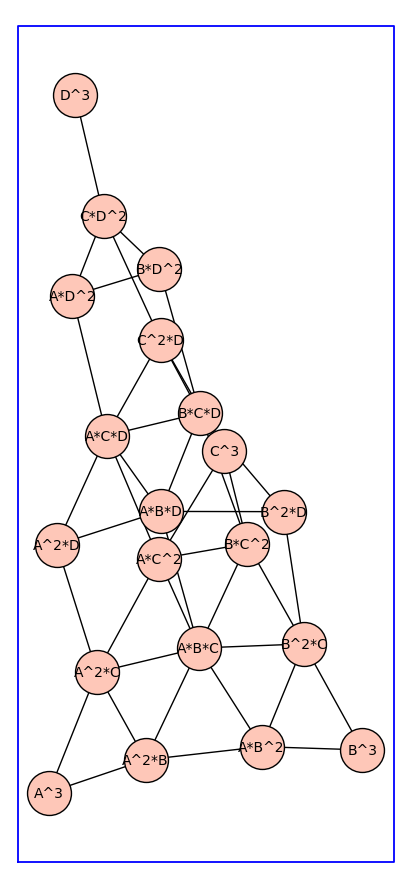
Next, we repeat the above example using a different choice of vertex labels.
H=add_vertex_monomials(G0,method='alpha')
H.show(figsize=4,graph_border=True)
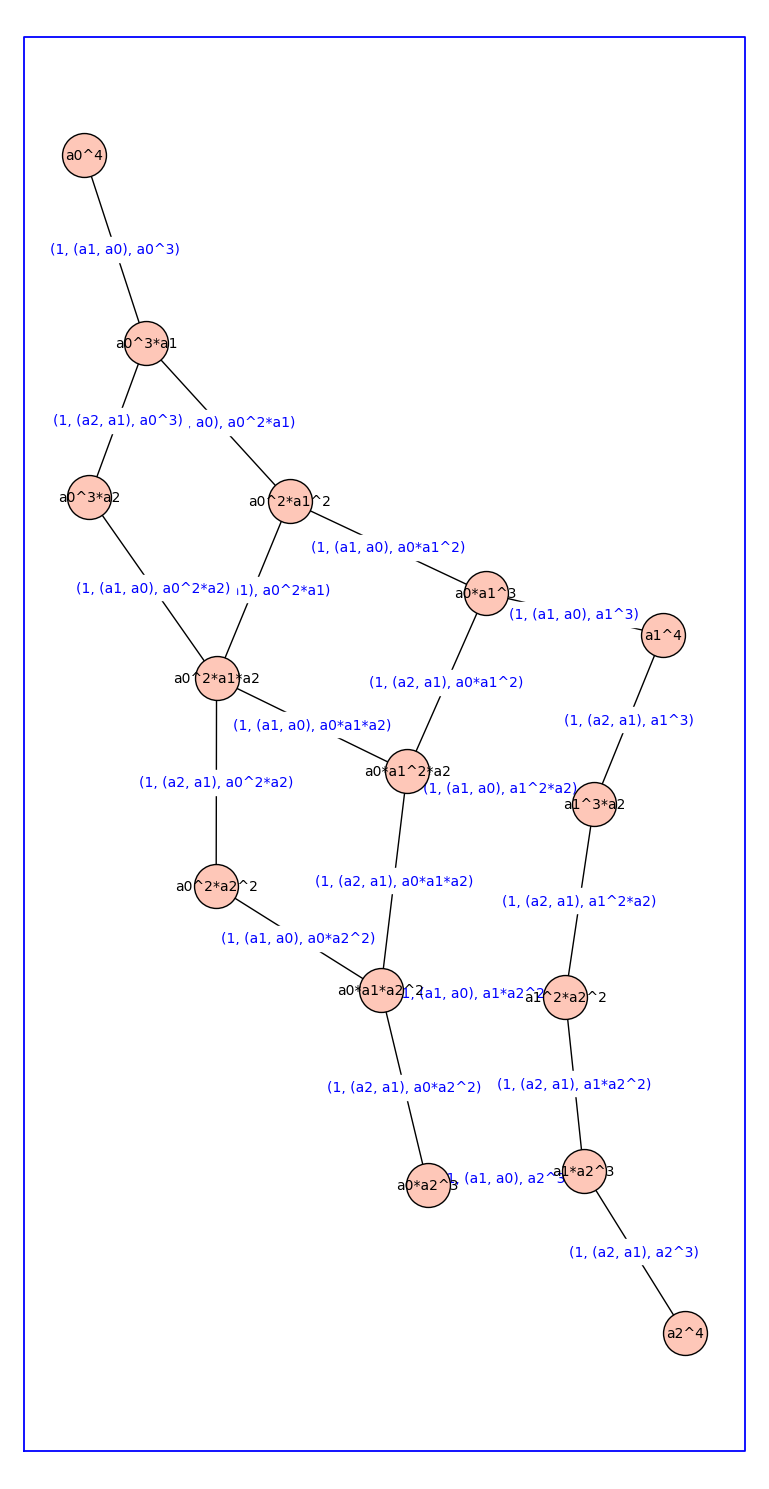
H3 = reduced_cartesian_power_ver2(H,3)
H3.show(figsize=12,graph_border=True,vertex_size=1000)
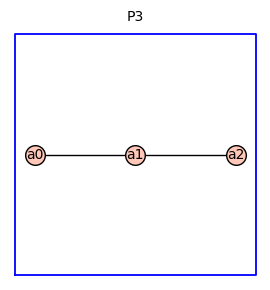
This next example shows the reduced graph power \(P_3^{(4)}\).
P3 = graphs.PathGraph(3)
P3 = add_vertex_monomials(P3)
P3.show(figsize=4,graph_border=True,title='P3')
P3_4 = reduced_cartesian_power_ver2(P3,4)
P3_4.show(figsize=20,graph_border=True,vertex_size=1000)
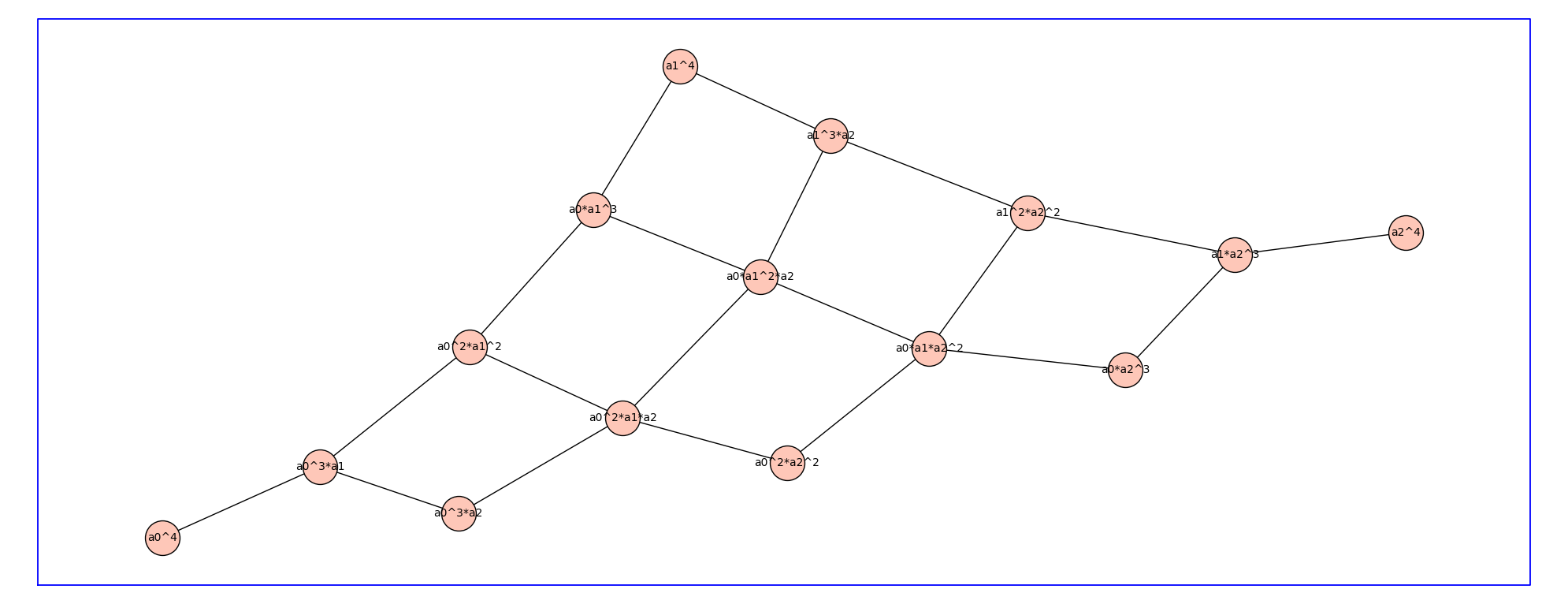
The edge labels are 3-tuples.
The first element is a combinatorial coefficient.
The second element is a tuple indicating the monomeric transition associated with the edge.
The third element is the context.
UNDER CONSTRUCTION#
The edge labels are not yet working. Do I need this? Should it be a directed graph?
P3_4.show(figsize=20,graph_border=True,vertex_size=1000,edge_labels=True)