States and transitions#
The process of receptor modeling often begins by specifying the molecular conformations (states) to be considered and the transitions between these states.
Undirected graphs as state-transition diagrams#
For example, the following graph may represent a receptor model that has four states:
G_undirected = Graph({1: [2, 3], 2: [3], 3: [4]})
vertex_positions = {1: (0, 0), 2: (1, 1.41), 3: (2, 0), 4: (4,0)}
G_undirected.plot(figsize=8,pos=vertex_positions,graph_border=True)
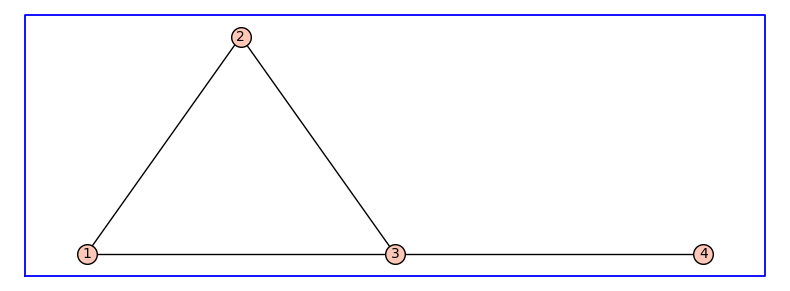
The graph G_undirected is constructed by calling the Sagemath command Graph() with a dictionary that associates neighbors to each vertex. The vertices of the graph G are the integers 1, 2, 3, and 4. The method plot() shows the graph G_undirected using a dictionary vertex_positions that specifies the locations of each vertex.
The adjacency matrix of this graph is
G_undirected.adjacency_matrix()
[0 1 1 0]
[1 0 1 0]
[1 1 0 1]
[0 0 1 0]
The elements of the adjacency matrix indicate whether pairs of vertices are adjacent or not in the graph. The adjacency matrix of a graph should be distinguished from its incidence matrix and its degree matrix.
Note
The graphs used here to represent receptor states and transitions will be both connected and simple.
Connected: at least one path joins every pair of vertices.
Simple: no loops or multiple edges.
Directed graphs as state-transition diagrams#
In the context of receptor modeling, the undirected graph above is interpreted as a short-hand for the following directed graph.
G_directed = G_undirected.to_directed()
G_directed.plot(figsize=8,edge_labels=True,pos=vertex_positions,graph_border=True)
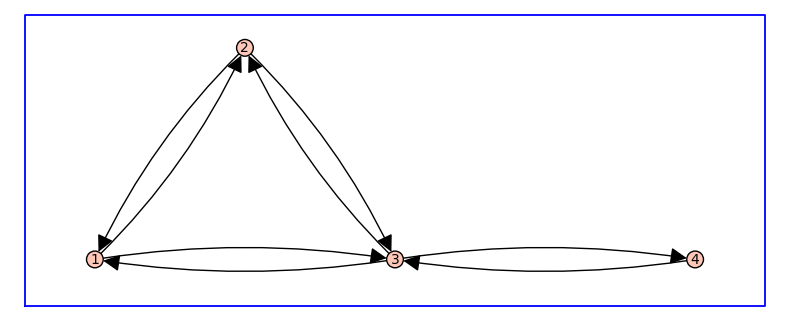
The method to_directed() produces G_directed as the symmetric digraph associated to G_undirected, in which adjacent vertices are connected in both directions.
Note
Receptor state-transition diagrams will always be symmetric directed graphs, that is, for every edge from vertex i to vertex j, there is also an edge from vertex j to vertex i. Thus, the state-transtion diagrams for a receptor model may, for simplicity, be illustrated as an undirected graph. We will refer to such undirected graphs as the topology of the receptor model.
Transition rate constants#
In the context of receptor modeling, state-transition diagrams are usually weighted, as shown here.
var('a12, a21, a13, a31, a23, a32, a34, a43')
d = {1: {2:a12, 3:a13}, 2: {1:a21, 3:a23}, 3: {2:a32, 1:a31, 4:a34}, 4: {3:a43}};
G = DiGraph(d,weighted=True)
G.plot(figsize=8,edge_labels=True,pos=vertex_positions,graph_border=True)
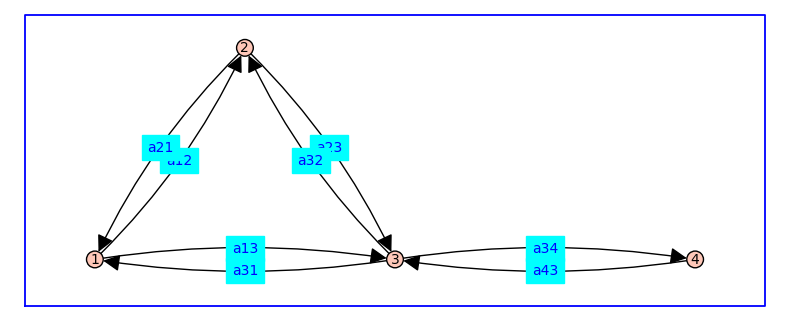
In the code above, a directed graph G is constructed by calling the Sagemath command DiGraph(). The input argument d is a Python dictionary that assigns out-neighbors to each vertex and corresponding edge labels.
The edge labels are not strings, but symbolic variables defined using Sagemath’s var command.
For example, the symbolic variable a12 stands for the rate of transition between state 1 and 2.
Because these rate constants are symbolic variables, Sagemath will evaluate expressions such as
f = a12 * (a21 + a13)^2 / a13
f.expand()
a12*a13 + 2*a12*a21 + a12*a21^2/a13
One reason for using symbolic variables is that we can produce symbolic expressions important quantities using the module for graphs and digraphs available in Sagemath. For example, the weighted adjacency matrix for G is
G.weighted_adjacency_matrix()
[ 0 a12 a13 0]
[a21 0 a23 0]
[a31 a32 0 a34]
[ 0 0 a43 0]
The Laplacian matrix of G is
G.laplacian_matrix()
[ a21 + a31 -a12 -a13 0]
[ -a21 a12 + a32 -a23 0]
[ -a31 -a32 a13 + a23 + a43 -a34]
[ 0 0 -a43 a34]
This matrix is sometimes referred to as the combinatorial Laplacian matrix of the weighted directed graph G.

