Plot Binding Curves#
%run MyTools.ipynb
import numpy as np
import random
from matplotlib import pyplot as plt
#G = graphs.HouseGraph()
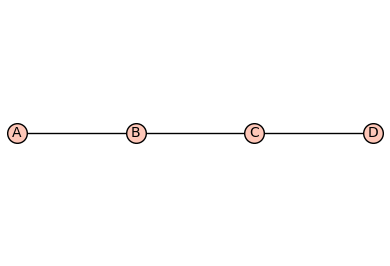
G = graphs.PathGraph(4)
(G, T, KappaEta, A) = enumerate_allosteric_parameters(G,method='alpha',verbose=False)
G.show(edge_labels=False)
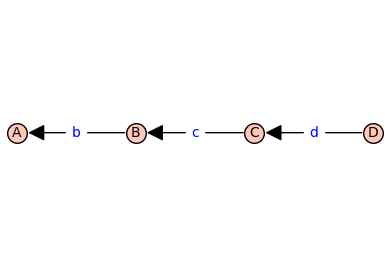
T.show(edge_labels=True)
show(table(KappaEta))
show(A)
mydoc(myfun, method='both')
Display the signature of myfun followed by the docstring of myfun.
Set method ='signature' or 'method = 'docstring' or 'method = 'both'
my_graph_show(G)
Show a graph Greg’s way.
Prints the vertices and edges of the graph G.
add_vertex_monomials(G=Graph on 0 vertices, method='integer', ring=False)
Add monomials to vertices of a graph.
The add_vertex_monomials function takes a graph G, as well as optional parameters method and ring. The function creates a new graph H with vertices labeled by monomials. The monomials are chosen based on the number of vertices in G. If the method parameter is set to ‘alpha’ and the number of vertices in G is less than or equal to 10, the monomials are chosen as alphabetical letters (‘a’ to ‘k’). Otherwise, the monomials are chosen as strings of the form ‘a0’, ‘a1’, …, ‘an-1’, where n is the number of vertices in G. The function then adds the vertices from G to H using the monomials as labels, and adds the edges from G to H using the monomials as endpoints. If the ring parameter is set to True, the function also creates a polynomial ring V with the chosen monomials and ‘invlex’ order, and returns both H and V. Otherwise, it returns only H.
INPUT:
G– graph object (default:Graph());method– integer (default:integer);
OUTPUT: The graph with monomials as vertices
EXAMPLES:
This example illustrates … ::
sage: A = ModuliSpace()
sage: A.point(2,3)
xxx
| \(1\) | \(2 \kappa_{\mathit{b}}\) | \(2 \kappa_{\mathit{b}} \kappa_{\mathit{c}}\) | \(2 \kappa_{\mathit{b}} \kappa_{\mathit{c}} \kappa_{\mathit{d}}\) |
| \(0\) | \(\kappa_{\mathit{b}}^{2} \eta_{\mathit{bb}}\) | \(2 \kappa_{\mathit{b}}^{2} \kappa_{\mathit{c}} \eta_{\mathit{bb}} \eta_{\mathit{bc}}\) | \(2 \kappa_{\mathit{b}}^{2} \kappa_{\mathit{c}} \kappa_{\mathit{d}} \eta_{\mathit{bb}} \eta_{\mathit{bc}} \eta_{\mathit{bd}}\) |
| \(0\) | \(0\) | \(\kappa_{\mathit{b}}^{2} \kappa_{\mathit{c}}^{2} \eta_{\mathit{bb}} \eta_{\mathit{bc}}^{2} \eta_{\mathit{cc}}\) | \(2 \kappa_{\mathit{b}}^{2} \kappa_{\mathit{c}}^{2} \kappa_{\mathit{d}} \eta_{\mathit{bb}} \eta_{\mathit{bc}}^{2} \eta_{\mathit{cc}} \eta_{\mathit{bd}} \eta_{\mathit{cd}}\) |
| \(0\) | \(0\) | \(0\) | \(\kappa_{\mathit{b}}^{2} \kappa_{\mathit{c}}^{2} \kappa_{\mathit{d}}^{2} \eta_{\mathit{bb}} \eta_{\mathit{bc}}^{2} \eta_{\mathit{cc}} \eta_{\mathit{bd}}^{2} \eta_{\mathit{cd}}^{2} \eta_{\mathit{dd}}\) |
n_states=G.order()
n_kappa=n_states-1
n_kappa_plus_eta = A.ngens()
n_eta=n_kappa_plus_eta-n_kappa
kappa_list = [A.gen(k) for k in range(n_kappa)]
print(f'kappa_list = {kappa_list}')
eta_list = [A.gen(k) for k in range(n_kappa,n_kappa_plus_eta)]
print(f'eta_list = {eta_list}')
phi=normalize(random_binary_array(n_states))
print(f'phi = {phi}')
Phi = dimerize(phi)
print(f'Phi = {Phi}')
print(f'KappaEta = {KappaEta}')
kappa_list = [kappa_b, kappa_c, kappa_d]
eta_list = [eta_bb, eta_bc, eta_cc, eta_bd, eta_cd, eta_dd]
phi = [0.5 0.5 0. 0. ]
Phi = [[np.float64(0.5), np.float64(0.25), np.float64(0.0), np.float64(0.0)], [0, np.float64(0.5), np.float64(0.0), np.float64(0.0)], [0, 0, np.float64(0.0), np.float64(0.0)], [0, 0, 0, np.float64(0.0)]]
KappaEta = [[1, 2*kappa_b, 2*kappa_b*kappa_c, 2*kappa_b*kappa_c*kappa_d], [0, kappa_b^2*eta_bb, 2*kappa_b^2*kappa_c*eta_bb*eta_bc, 2*kappa_b^2*kappa_c*kappa_d*eta_bb*eta_bc*eta_bd], [0, 0, kappa_b^2*kappa_c^2*eta_bb*eta_bc^2*eta_cc, 2*kappa_b^2*kappa_c^2*kappa_d*eta_bb*eta_bc^2*eta_cc*eta_bd*eta_cd], [0, 0, 0, kappa_b^2*kappa_c^2*kappa_d^2*eta_bb*eta_bc^2*eta_cc*eta_bd^2*eta_cd^2*eta_dd]]
# create the symbolic binding curve
FBindingKappaEta=make_symbolic_dimer_binding_curve(Phi,KappaEta)
show(FBindingKappaEta)
# add ligand dependencies randomly or manually
manual = False
var('X')
if manual:
d_kappa_X = dict({kappa_b: kappa_b*X, kappa_c: kappa_c*X})
else:
def Round_To_n(x, n):
return round(x, -int(np.floor(np.sign(x) * np.log10(abs(x)))) + n)
d_kappa_X=dict()
num_X = 0
while num_X == 0: # make sure there is at least one ligand binding edge (kappa*X)
for kappa in kappa_list:
k=Round_To_n(np.random.exponential(scale=1.0),2)
ligand_prob = 0.4
if np.random.random() <= ligand_prob:
d_kappa_X[kappa]=k*X
num_X += 1
else:
d_kappa_X[kappa]=k
print(d_kappa_X)
FBindingEtaX=FBindingKappaEta.subs(d_kappa_X)
show(FBindingEtaX)
{kappa_b: 0.928, kappa_c: 1.31*X, kappa_d: 0.437}
FastCallableFBindingEtaX = fast_callable(FBindingEtaX, vars=['X']+eta_list)
from scipy.optimize import minimize
import random
from matplotlib import pyplot as plt
# construct target function
# xrange and values x for data, x0 for smooth plots
xlogmin=-3
xlogmax=3
x = np.logspace(xlogmin,xlogmax,20)
x0 = np.logspace(xlogmin,xlogmax,1000)
# choose random eta's with 50% change of being 1 (unused)
eta_prob_one = 0.5
eta = np.random.exponential(scale=1.0,size=n_eta)
for i in range(n_eta):
if np.random.random() <= eta_prob_one:
eta[i]=1.0
y = FastCallableFBindingEtaX(x,*eta) + np.random.normal(size=x.size, scale=0.005) # w/ noise
y0 = FastCallableFBindingEtaX(x0,*eta)
# create hypercube and flip left/right
Q = graphs.CubeGraph(n_eta, embedding=2)
pos = Q.get_pos()
for v in Q.vertices(sort=False):
xpos,ypos = pos[v] # do not use x,y here! These are needed below
pos[v]=(n_eta-xpos,ypos)
# pos[v]=(ypos,xpos) this didn't look good
#Q.show(figsize=6,edge_labels=False,vertex_labels=True, vertex_size=100,edge_thickness=0.5,vertex_colors='white')
root = "1" * n_eta
d_level = dict()
for level in range(n_eta+1):
for v in Q.vertices(sort=True):
if level == Q.distance(v,root):
d_level[v]=level
print(d_level)
{'111111': 0, '011111': 1, '101111': 1, '110111': 1, '111011': 1, '111101': 1, '111110': 1, '001111': 2, '010111': 2, '011011': 2, '011101': 2, '011110': 2, '100111': 2, '101011': 2, '101101': 2, '101110': 2, '110011': 2, '110101': 2, '110110': 2, '111001': 2, '111010': 2, '111100': 2, '000111': 3, '001011': 3, '001101': 3, '001110': 3, '010011': 3, '010101': 3, '010110': 3, '011001': 3, '011010': 3, '011100': 3, '100011': 3, '100101': 3, '100110': 3, '101001': 3, '101010': 3, '101100': 3, '110001': 3, '110010': 3, '110100': 3, '111000': 3, '000011': 4, '000101': 4, '000110': 4, '001001': 4, '001010': 4, '001100': 4, '010001': 4, '010010': 4, '010100': 4, '011000': 4, '100001': 4, '100010': 4, '100100': 4, '101000': 4, '110000': 4, '000001': 5, '000010': 5, '000100': 5, '001000': 5, '010000': 5, '100000': 5, '000000': 6}
def do_eta_fit(eta_list):
# Define the objective function
def objective_function(params):
eta = params
residuals = y - FastCallableFBindingEtaX(x,*eta)
return np.sum(residuals ** 2)
# Initial guess for parameters
initial_guess = np.ones(n_eta)
# this loop makes elements of the tuple bnds
bnds = ()
k = 0 # nparams
for i in range(n_eta):
if eta_list[i]==1.0:
bnds += ((1, 1),) # eta = 1, not used
else:
bnds += ((0, None),) # eta non-negative
k += 1
# Perform optimization
result = minimize(objective_function, initial_guess, bounds=bnds)
# Extract and print optimized eta's
optimal_eta = result.x
np.set_printoptions(precision=2)
print( optimal_eta )
#print(f'{optimal_eta=}')
# Evaluate result
y0_result = FastCallableFBindingEtaX(x0,*optimal_eta)
res = y - FastCallableFBindingEtaX(x,*optimal_eta)
ssr = np.sum(res ** 2)
n = len(y)
aic = 2*(k+2)+n*np.log(ssr/(n-k))
# Plot result
plot_results = False
if plot_results:
plt.semilogx(x0,y0,linewidth=4,color='yellow',label='exact')
plt.semilogx(x,y,'+',color='black',label='sim data')
plt.semilogx(x0,y0_result,color='red',linestyle='dashed',label='fit')
np.set_printoptions(precision=2)
plt.title(f'aic={aic=},\ntarget eta={eta},\noptimal eta={optimal_eta}')
plt.xlabel('x')
plt.ylabel('phi',rotation = 0)
plt.legend()
plt.grid()
plt.show()
return aic
# make arcs point rightward to "lower" levels only (fewer parameters used, more 1's)
Q_directed = Q.to_directed()
for e in Q_directed.edges(sort=False):
if d_level[e[0]]<d_level[e[1]]:
Q_directed.delete_edge(e)
#show(Q_directed)
max_depth = 4
d_aic = dict();
Q_aic = copy(Q_directed)
for level in range(max_depth+1,n_eta+1): # delete levels not used
#print(f'{max_depth=} so deleting {level=}')
for vert, lev in d_level.items():
if lev == level:
Q_aic.delete_vertex(vert)
for level in range(0,max_depth+1):
for vert, lev in d_level.items():
if lev == level:
if Q_aic.has_vertex(vert):
#print(type(vert))
aic = do_eta_fit([int(char) for char in vert])
d_aic[vert]=str(f'{aic:.3f}')
Q_aic.set_vertex(vert,aic)
#print(f"level {lev} : eta_mask={vert} {aic=:.2f}")
hang = 0
for nv in Q_aic.neighbors_out(vert):
#print(f'looking at vert {vert} --> {nv}')
if float(d_aic[vert])>=float(d_aic[nv]): # float from str
#print(f'deleted edge {vert=} {d_aic[vert]=} -> {nv=} {d_aic[nv]=}')
Q_aic.delete_edge(vert,nv)
else:
hang+=1
if hang==0 and vert != root:
Q_aic.delete_vertex(vert)
[1. 1. 1. 1. 1. 1.]
[0.48 1. 1. 1. 1. 1. ]
[1. 0.45 1. 1. 1. 1. ]
[1.00e+00 1.00e+00 4.51e-17 1.00e+00 1.00e+00 1.00e+00]
[1. 1. 1. 0. 1. 1.]
[1. 1. 1. 1. 0. 1.]
[1.00e+00 1.00e+00 1.00e+00 1.00e+00 1.00e+00 2.23e-17]
[0.93 0.45 1. 1. 1. 1. ]
[0.97 1. 0. 1. 1. 1. ]
[0.85 1. 1. 0. 1. 1. ]
[0.71 1. 1. 1. 0. 1. ]
[0.52 1. 1. 1. 1. 0. ]
[1. 0.52 0.55 1. 1. 1. ]
[1. 0.46 1. 0.96 1. 1. ]
[1. 0.52 1. 1. 0.16 1. ]
[1. 0.45 1. 1. 1. 1. ]
[1. 1. 0.11 0. 1. 1. ]
[1. 1. 0. 1. 0.43 1. ]
[1.00e+00 1.00e+00 2.19e-17 1.00e+00 1.00e+00 9.08e-01]
[1. 1. 1. 0. 0.62 1. ]
[1. 1. 1. 0. 1. 1.1]
[1. 1. 1. 1. 0. 1.04]
[0.95 0.49 0.67 1. 1. 1. ]
[0.93 0.46 1. 0.94 1. 1. ]
[0.95 0.49 1. 1. 0.41 1. ]
[0.93 0.45 1. 1. 1. 1. ]
[1.01 1. 0.11 0. 1. 1. ]
[0.97 1. 0. 1. 0.5 1. ]
[0.97 1. 0. 1. 1. 0.92]
[0.85 1. 1. 0. 0.57 1. ]
[0.85 1. 1. 0. 1. 0.97]
[0.71 1. 1. 1. 0. 0.91]
[1. 0.52 0.55 0.96 1. 1. ]
[1. 0.52 0.64 1. 0.78 1. ]
[1. 0.52 0.56 1. 1. 0.97]
[1. 0.52 1. 0.98 0.14 1. ]
[1. 0.46 1. 0.96 1. 1. ]
[1. 0.52 1. 1. 0.16 0.87]
[1. 1. 0.11 0. 0.82 1. ]
[1. 1. 0.11 0. 1. 1.11]
[1. 1. 0. 1. 0.43 1.02]
[1. 1. 1. 0. 0.61 1.05]
[0.95 0.51 0.67 0.89 1. 1. ]
[0.95 0.49 0.73 1. 0.86 1. ]
[0.95 0.49 0.67 1. 1. 0.98]
[0.95 0.5 1. 0.98 0.4 1. ]
[0.93 0.46 1. 0.94 1. 1. ]
[0.95 0.49 1. 1. 0.41 0.92]
[1.01 1. 0.11 0. 0.7 1. ]
[1.01 1. 0.11 0. 1. 0.98]
[0.97 1. 0. 1. 0.5 0.96]
[0.85 1. 1. 0. 0.57 1. ]
[1. 0.53 0.62 0.9 0.83 1. ]
[1. 0.52 0.55 0.96 1. 0.97]
[1. 0.52 0.64 1. 0.78 0.97]
[1. 0.52 1. 0.97 0.14 0.88]
[1. 1. 0.11 0. 0.81 1.05]
print(d_aic)
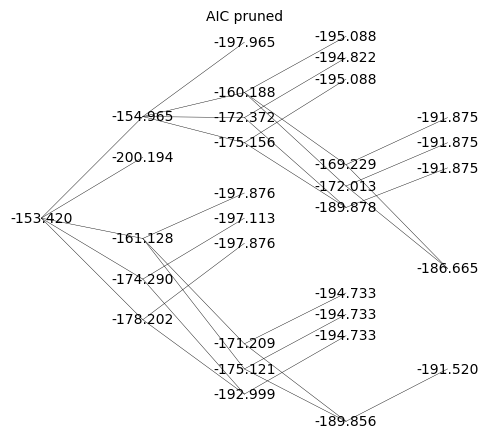
{'111111': '-153.420', '011111': '-154.965', '101111': '-200.194', '110111': '-178.202', '111011': '-174.290', '111101': '-161.128', '111110': '-151.928', '001111': '-197.965', '010111': '-175.156', '011011': '-172.372', '011101': '-160.188', '011110': '-152.984', '100111': '-197.876', '101011': '-197.113', '101101': '-197.876', '101110': '-197.112', '110011': '-192.999', '110101': '-175.121', '110110': '-175.121', '111001': '-171.209', '111010': '-171.209', '111100': '-158.047', '000111': '-195.088', '001011': '-194.822', '001101': '-195.088', '001110': '-194.823', '010011': '-189.878', '010101': '-172.013', '010110': '-172.013', '011001': '-169.229', '011010': '-169.229', '011100': '-157.045', '100011': '-194.733', '100101': '-194.733', '100110': '-194.733', '101001': '-194.733', '101010': '-193.969', '101100': '-194.733', '110001': '-189.856', '110010': '-189.856', '110100': '-171.978', '111000': '-168.065', '000011': '-191.875', '000101': '-191.875', '000110': '-191.875', '001001': '-191.875', '001010': '-191.610', '001100': '-191.875', '010001': '-186.665', '010010': '-186.665', '010100': '-168.800', '011000': '-166.016', '100001': '-191.520', '100010': '-191.520', '100100': '-191.520', '101000': '-191.520', '110000': '-186.643'}
Q_aic_undirected = Q_aic.to_undirected()
Q_aic_undirected.show(figsize=6,edge_labels=False,vertex_labels=d_aic, vertex_size=0,edge_thickness=0.3,vertex_colors='white',title='AIC pruned')
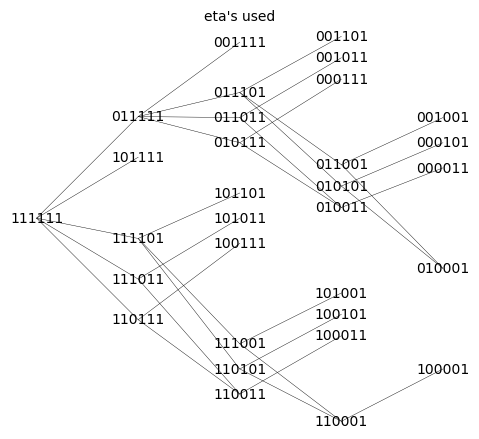
Q_aic_undirected.show(figsize=6,edge_labels=False, vertex_size=0,edge_thickness=0.3,vertex_colors='white',title='eta\'s used')